
Christian BRAENDLE
Gene-environment interactions in development and evolution
Main interests
- Phenotypic plasticity, genotype-by-environment interactions and genetic assimilation
- Genotype-phenotype map: from developmental variation to life history variation
- Evolution and ecology of Caenorhabditis nematodes
Scientific Questions
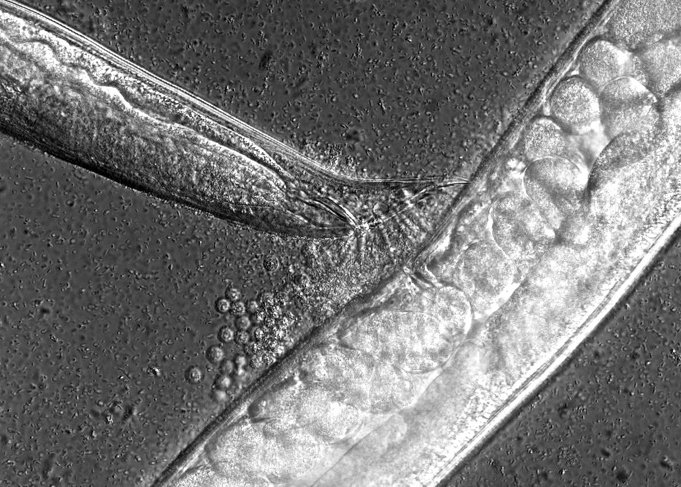
Environmental variation is a key force shaping the function and evolution of developmental mechanisms. Therefore, it is essential to understand how environmental variation modulates developmental processes and resulting phenotypic variation. Much of our research therefore addresses the questions of how developmental systems respond to environmental variation and how such responses evolve. Using the nematode Caenorhabditis elegans and related species, we aim to answer four main questions: How does development integrate variable environmental information to alter phenotypic outcomes? How do developmental responses to environmental variation evolve? How do quantiative developmental traits evolve within species? What are developmental changes underlying adaptive evolution of life histories? We investigate these questions by focusing on molecularly well-characterized processes, such as vulval cell fate patterning, germline development and dauer formation.
Our Strategy
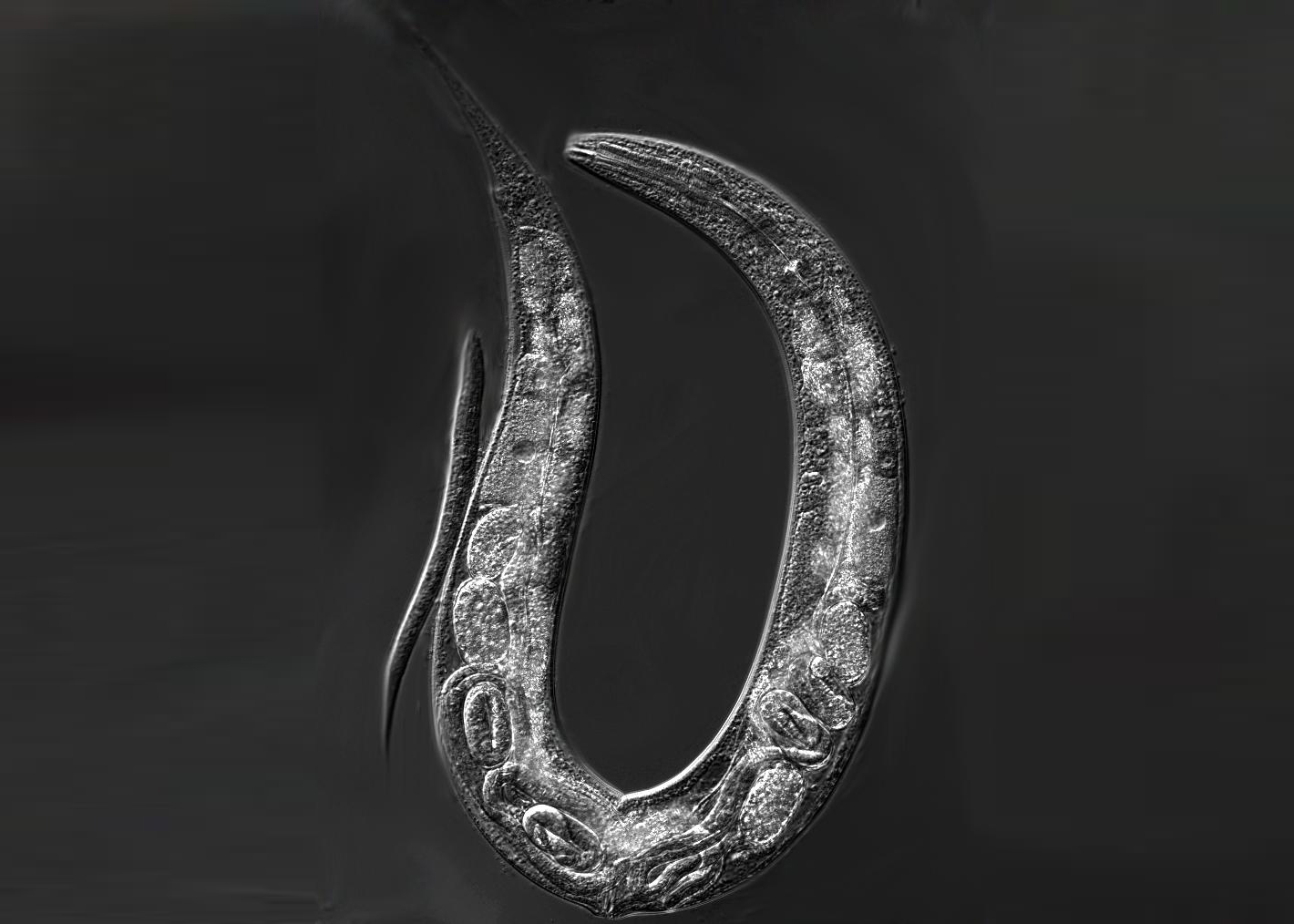
Although genotype-by-environment interactions are common and important determinants of phenotypic variation, the mechanisms by which genetic and environmental variation interact to generate trait variation remain poorly understood. Our projects thus aim to characterize the molecular and developmental basis of genotype-by-environment interactions, how such interactions evolve and how they in turn may impact the evolutionary process itself. In our research, we use the nematode Caenorhabditis elegans and related species as model organisms, and we integrate quantitative experimental approaches from developmental and evolutionary genetics.
Research Aims
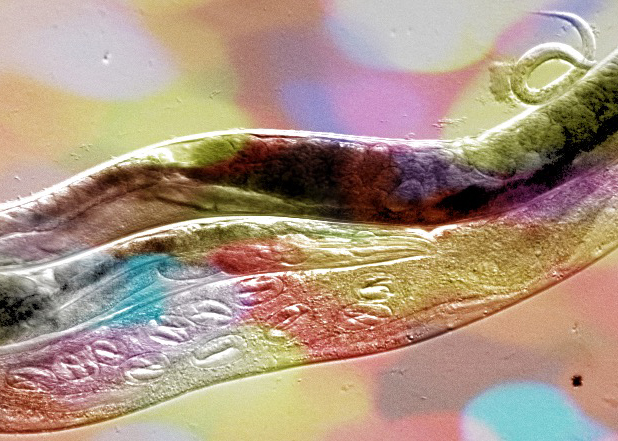
Phenotypic plasticity, genotype-by-environment interactions and genetic assimilation
We focus on the environmental context-dependence of developmental processes (e.g. germ cell proliferation, gametogenesis, dauer formation) to characterize the molecular basis and evolution of phenotypic plasticity. Primarily, we aim to identify developmental and molecular determinants of natural variation in these phenotypes through a combination of quantitative and developmental genetic approaches.
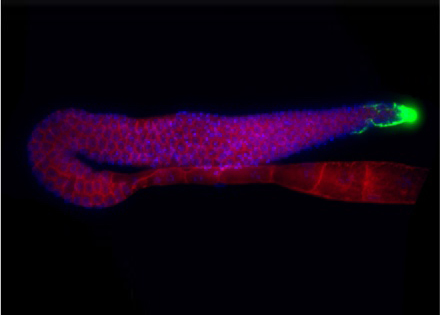
Genotype-phenotype map: from developmental variation to life history variation
How complex traits, such as life history traits, emerge through developmental integration of numerous gene-environment, gene-gene interactions, and higher-order interactions among cells and tissues, remains largely unknown. We therefore study the nematode hermaphrodite germline to ask how variation in different parameters of this developmental system translates into variation in reproductive life histories.
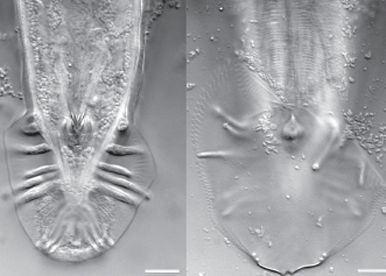
Evolution and ecology of Caenorhabditis nematodes
Although C. elegans is increasingly being used in evolutionary studies, there is still very little information on its natural history, ecology, phylogenetic context, and the genetic structure of its natural populations. We therefore contribute to current community efforts to sample and characterize natural Caenorhabditis populations to generate a more comprehensive evolutionary ecological context for C. elegans and its close relatives.
Researchers
 GIMOND Clotilde - +33 489150842
GIMOND Clotilde - +33 489150842
Postdocs
 GONZALEZ SOMERMEYER Louisa - +33 489150842
GONZALEZ SOMERMEYER Louisa - +33 489150842
PreDocs
 SANDJAK Asma - +33 489150842
SANDJAK Asma - +33 489150842 LE PARC Amélie - +33 489150842
LE PARC Amélie - +33 489150842
Engineers & Technicians
 BOUVET Orianne - +33 489150842
BOUVET Orianne - +33 489150842 LEROUX Delphine - +33 489150842
LEROUX Delphine - +33 489150842 BOYER Salomé - +33 489150842
BOYER Salomé - +33 489150842 GOMES SANCHES VAZ Eliane - +33 489150842
GOMES SANCHES VAZ Eliane - +33 489150842 VENNIN Tom - +33 489150842
VENNIN Tom - +33 489150842
Recent publications
- Greenway, R, Dalan, L, Braendle, C, Félix, MA, Ding, SS. Differential Phoretic Vector Use Among Sympatric Caenorhabditis Nematodes and an Association With Invasive Nitidulid Beetles in Southwestern Germany. Ecol Evol. 2026;16 :e73510. doi: 10.1002/ece3.73510. PubMed PMID:42088392 PubMed Central PMC13138117.
- Wang, B, Moya, ND, Tanny, RE, Sauria, MEG, O'Connor, LM, Khorshidian, A et al.. Global genomic diversity of the selfing nematode Caenorhabditis tropicalis correlates with geography. bioRxiv. 2026; :. doi: 10.64898/2026.04.05.716573. PubMed PMID:41993510 PubMed Central PMC13081884.
- Crombie, TA, Mckeown, R, Widmayer, SJ, Shaver, AO, Moya, ND, Collins, JB et al.. Natural variation suggests candidate genes underlying Caenorhabditis elegans susceptibility to diverse toxicants. Toxicol Sci. 2026;209 (3):. doi: 10.1093/toxsci/kfag019. PubMed PMID:41706139 .
- Moya, ND, Wang, B, Tanny, RE, Sauria, MEG, O'Connor, LM, Khorshidian, A et al.. Caenorhabditis briggsae ancestral genomic hyper-diversity contrasts with globally distributed genome-wide haplotypes. bioRxiv. 2025; :. doi: 10.64898/2025.12.08.693002. PubMed PMID:41427407 PubMed Central PMC12715550.
- Crombie, TA, McKeown, R, Widmayer, SJ, Shaver, AO, Moya, ND, Collins, JB et al.. Natural variation suggests candidate genes underlying Caenorhabditis elegans susceptibility to diverse toxicants. bioRxiv. 2025; :. doi: 10.1101/2025.11.30.691394. PubMed PMID:41377522 PubMed Central PMC12687771.
- Gimond, C, Poullet, N, Vielle, A, Demoinet, E, Braendle, C. Developmental plasticity of hermaphrodite sperm production across environments in Caenorhabditis elegans. PLoS One. 2025;20 (11):e0336162. doi: 10.1371/journal.pone.0336162. PubMed PMID:41202014 PubMed Central PMC12594432.
- Picao-Osorio, J, Bouleau, C, Gonzalez de la Rosa, PM, Stevens, L, Fekonja, N, Blaxter, M et al.. Evolution of developmental bias explains divergent patterns of phenotypic evolution in two nematode clades. Proc Natl Acad Sci U S A. 2025;122 (34):e2507529122. doi: 10.1073/pnas.2507529122. PubMed PMID:40828025 PubMed Central PMC12403097.
- Vigne, P, Braendle, C. Natural variation in cold and heat survival among temperate and tropical Caenorhabditis briggsae. MicroPubl Biol. 2025;2025 :. doi: 10.17912/micropub.biology.001645. PubMed PMID:40636101 PubMed Central PMC12238878.
- Braendle, C, Paaby, A. Life history in Caenorhabditis elegans: from molecular genetics to evolutionary ecology. Genetics. 2024;228 (3):. doi: 10.1093/genetics/iyae151. PubMed PMID:39422376 PubMed Central PMC11538407.
- Mignerot, L, Gimond, C, Bolelli, L, Bouleau, C, Sandjak, A, Boulin, T et al.. Natural variation in the Caenorhabditis elegans egg-laying circuit modulates an intergenerational fitness trade-off. Elife. 2024;12 :. doi: 10.7554/eLife.88253. PubMed PMID:38564369 PubMed Central PMC10987095.
- Fausett, SR, Laury, CA, Magallon, RE, Braendle, C. Simplified Quantification of Progenitor Zone Size, an Indicator of Germ Stem Cell Niche Activity, in the Nematode Caenorhabditis elegans. Methods Mol Biol. 2025;2939 :183-191. doi: 10.1007/7651_2024_536. PubMed PMID:38507211 .
- Fausett, SR, Sandjak, A, Billard, B, Braendle, C. Higher-order epistasis shapes natural variation in germ stem cell niche activity. Nat Commun. 2023;14 (1):2824. doi: 10.1038/s41467-023-38527-0. PubMed PMID:37198172 PubMed Central PMC10192456.
- Huang, Y, Lo, YH, Hsu, JC, Le, TS, Yang, FJ, Chang, T et al.. Widespread sex ratio polymorphism in Caenorhabditis nematodes. R Soc Open Sci. 2023;10 (3):221636. doi: 10.1098/rsos.221636. PubMed PMID:36938539 PubMed Central PMC10014251.
- Gimond, C, Poullet, N, Braendle, C. Isolating Caenorhabditis elegans from the Natural Habitat. Methods Mol Biol. 2022;2468 :283-292. doi: 10.1007/978-1-0716-2181-3_15. PubMed PMID:35320571 .
- Fausett, S, Poullet, N, Gimond, C, Vielle, A, Bellone, M, Braendle, C et al.. Germ cell apoptosis is critical to maintain Caenorhabditis elegans offspring viability in stressful environments. PLoS One. 2021;16 (12):e0260573. doi: 10.1371/journal.pone.0260573. PubMed PMID:34879088 PubMed Central PMC8654231.
- Grover, M, Fasseas, MK, Essmann, C, Liu, K, Braendle, C, Félix, MA et al.. Infection of C. elegans by Haptoglossa Species Reveals Shared Features in the Host Response to Oomycete Detection. Front Cell Infect Microbiol. 2021;11 :733094. doi: 10.3389/fcimb.2021.733094. PubMed PMID:34722333 PubMed Central PMC8552708.
- Lee, D, Zdraljevic, S, Stevens, L, Wang, Y, Tanny, RE, Crombie, TA et al.. Balancing selection maintains hyper-divergent haplotypes in Caenorhabditis elegans. Nat Ecol Evol. 2021;5 (6):794-807. doi: 10.1038/s41559-021-01435-x. PubMed PMID:33820969 PubMed Central PMC8202730.
- Vigne, P, Gimond, C, Ferrari, C, Vielle, A, Hallin, J, Pino-Querido, A et al.. A single-nucleotide change underlies the genetic assimilation of a plastic trait. Sci Adv. 2021;7 (6):. doi: 10.1126/sciadv.abd9941. PubMed PMID:33536214 PubMed Central PMC7857674.
- Noble, LM, Yuen, J, Stevens, L, Moya, N, Persaud, R, Moscatelli, M et al.. Selfing is the safest sex for Caenorhabditis tropicalis. Elife. 2021;10 :. doi: 10.7554/eLife.62587. PubMed PMID:33427200 PubMed Central PMC7853720.
- Ben-David, E, Pliota, P, Widen, SA, Koreshova, A, Lemus-Vergara, T, Verpukhovskiy, P et al.. Ubiquitous Selfish Toxin-Antidote Elements in Caenorhabditis Species. Curr Biol. 2021;31 (5):990-1001.e5. doi: 10.1016/j.cub.2020.12.013. PubMed PMID:33417886 .
2012 - Fellow at the Institute for Advanced Study, Berlin
2010 - Schlumberger Award
2008 - ATIP, CNRS
1995 - Fund for Young Talented People, Swiss Study Foundation
1994 - Winner of highest award in the Swiss science contest for young people “Schweizer Jugend Forscht”

Christian Braendle awarded HFSP research grant for groundbreaking work on adaptation to extreme environments
Read More
Maternal self-sacrifice in C. elegans: an example of genetic assimilation
Read More
Evolution of a trade-off between two life history traits in the nematode C. elegans.
Read More

When iBV researchers meet Art at the Museum of Modern Art
Read More

Genetic basis of sperm size in C. elegans: a role for the chromatin remodeling complex NURF-1
Read More
iBV - Institut de Biologie Valrose
"Sciences Naturelles"
Université Nice Sophia Antipolis
Faculté des Sciences
Parc Valrose
06108 Nice cedex 2
