
Olivier SORIANI
Regulation of ion channels in cancer
Main interests
- Understanding how ion channels participate to the functional heterogeneity of the cancer tissue
- Understanding the role of ion channels in the dysregulation of cancer-associated signaling pathways
- Finding ion channel-associated targets to reduce tumour invasiveness
- Defining the physiopathological mechanisms associated to ion channel mutations in red blood cells
Scientific Questions
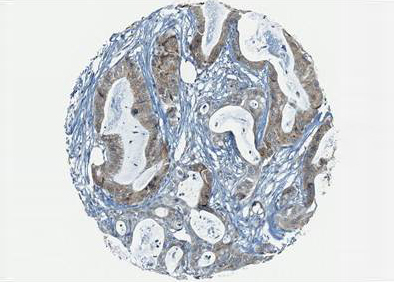
Ion channels are membrane proteins involved in many physiological and pathophysiological processes. Our group has focused his research on the role of these membrane proteins in cancer and hereditary hemolytic anemias.
1) Ion channel and cancer: Ion channels are involved in coupling extracellular events to cell behaviour. They contribute to the hallmarks of cancers and clinical outcome. Therefore, it is crucial to gain knowledge on the oncogenic mechanisms behind ion channel dysregulation to enhance current treatments.
2) Hereditary stomatocytosis is responsible for hemolytic anemia that is caused by mutations in red blood cell ion channels. Understanding ion channel alterations behing mutations in patients is a prerequisite to treat this rare disease.
Our Strategy
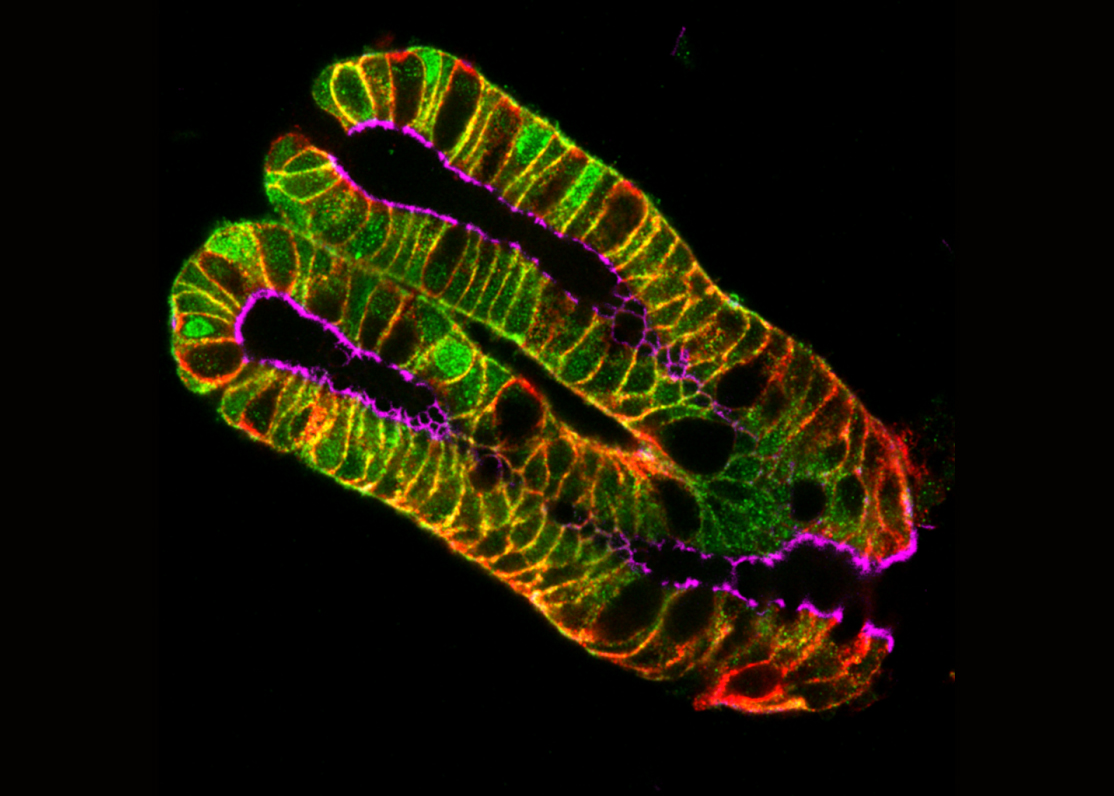
1) Using 3D cell culture, co-culture with cell lines specifically expressing or invalidated for targetted ion channels and their regulators, we study the impact of various components of the tumour micro environmment (TME) (such as cancer fibroblast-derived matrix) on tumour cell electrical activity, as well as the link between ion channel activity and tumour cell behavior in vitro and in vivo. This led to the characterization of several ion channels and regulatory proteins as potential players in TME-induced cancer cell invasiveness.
2) By organizing a partnership with Hematologists, we characterize the different mutations identified in patients suffering hereditary stomatocytosis. The moleculars features of mutated ion channels are studied as well as their effect on red blood cell homeostasis.
Research Aims
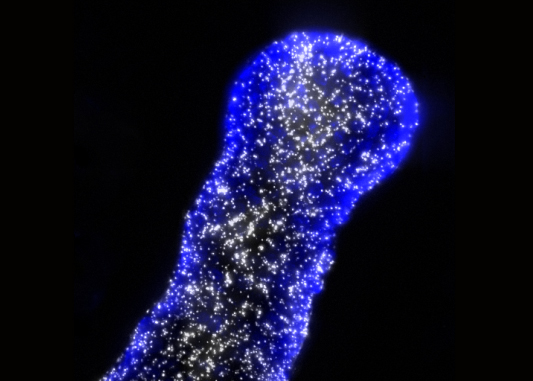
In the last few years, the study of the tumour microenvironment has taken a significant place in the field of cancer research. We have shown that SigmaR1, a stress-activated ion channel chaperone, plays a key function by controlling ion channels in response to TME. Therefore, We aim at characterizing the network of ion channels controlled by SigmaR1 in cancer cells and its functions in metastasic process in vivo and in vitro.
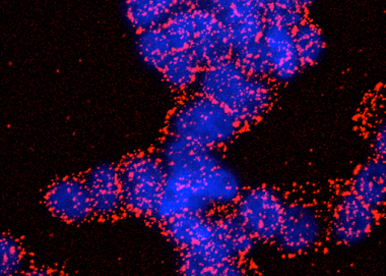
SigmaR1 is considered as a potential pharmacological target in neurodegenerative diseases and cancer. However, the molecular mechanisms underlying the coupling between SigmaR1 and ion channels need to be characterized to develop drugs modulating this interaction. By combining electrophysiology, biochemistry and imagery we aim at defining the dynamics of interaction between the partners, and the way sigma ligands interfere with SigmaR1/ion channel complexes.
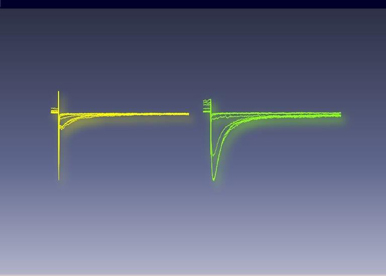
At the moment, two different genes have been linked to the dehydrated form of hereditary stomatocytosis: KCNN4 coding for the Gardos channel (our work) and Piezo1 coding for a mechano-sensitive ion channel. We aim at a better understanding of the connection between these two ion channels to develop treatments for this pathology as well as for other red blood cell pahtologies where volume homeostasis is impaired.
Researchers
 GUIZOUARN Hélène - +33 489150812
GUIZOUARN Hélène - +33 489150812 BORGESE Franck - +33 489150812
BORGESE Franck - +33 489150812 RAPETTI-MAUSS Raphaël - +33 489150813
RAPETTI-MAUSS Raphaël - +33 489150813
PreDocs
 TORRES ORTIZ Estefania - +33 489150812
TORRES ORTIZ Estefania - +33 489150812
Engineers & Technicians
 SOUSA DA VEIGA Anette - +33 R
SOUSA DA VEIGA Anette - +33 R RIPOCHE Léa - +33 489150812
RIPOCHE Léa - +33 489150812
Students
 MISSOUM Youssra - +33 489150812
MISSOUM Youssra - +33 489150812  RGUILI Samy - +33 489150812
RGUILI Samy - +33 489150812  HERANT Yohann - +33 R
HERANT Yohann - +33 R
Masters
 COLIN Clément - +33 489150812
COLIN Clément - +33 489150812  RGUILI Samy - +33 R
RGUILI Samy - +33 R
- Berenguier, C, Chen, X, Allegrini, B, Guizouarn, H, Borgese, F, Etchebest, C et al.. Cancer-associated loss-of-function mutations in KCNQ1 enhance Wnt/β-catenin signalling disrupting epithelial homeostasis. Oncogene. 2025;44 (31):2715-2729. doi: 10.1038/s41388-025-03447-4. PubMed PMID:40410368 .
- Allegrini, B, Mignotet, M, Rapetti-Mauss, R, Borgese, F, Soriani, O, Guizouarn, H et al.. A new regulation mechanism for KCNN4, the Ca2+-dependent K+ channel, by molecular interactions with the Ca2+pump PMCA4b. J Biol Chem. 2025;301 (2):108114. doi: 10.1016/j.jbc.2024.108114. PubMed PMID:39716493 PubMed Central PMC11787511.
- Pasquier, C, Guerlais, V, Pallez, D, Rapetti-Mauss, R, Soriani, O. A network embedding approach to identify active modules in biological interaction networks. Life Sci Alliance. 2023;6 (9):. doi: 10.26508/lsa.202201550. PubMed PMID:37339804 PubMed Central PMC10282331.
- Rapetti-Mauss, R, Nigri, J, Berenguier, C, Finetti, P, Tubiana, SS, Labrum, B et al.. SK2 channels set a signalling hub bolstering CAF-triggered tumourigenic processes in pancreatic cancer. Gut. 2023;72 (4):722-735. doi: 10.1136/gutjnl-2021-326610. PubMed PMID:36882214 .
- Cannone, S, Greco, MR, Carvalho, TMA, Guizouarn, H, Soriani, O, Di Molfetta, D et al.. Cancer Associated Fibroblast (CAF) Regulation of PDAC Parenchymal (CPC) and CSC Phenotypes Is Modulated by ECM Composition. Cancers (Basel). 2022;14 (15):. doi: 10.3390/cancers14153737. PubMed PMID:35954400 PubMed Central PMC9367491.
- Braud, V, Cazals, F, Duca, M, Mus-Veteau, I, Pasquier, C, Soriani, O et al.. [Highlights of the UCAncer 2021 symposium on multidisciplinary approaches to cancer research]. Bull Cancer. 2022;109 (4):505-509. doi: 10.1016/j.bulcan.2022.02.002. PubMed PMID:35277240 .
- Yamaguchi, Y, Allegrini, B, Rapetti-Mauss, R, Picard, V, Garçon, L, Kohl, P et al.. Hereditary Xerocytosis: Differential Behavior of PIEZO1 Mutations in the N-Terminal Extracellular Domain Between Red Blood Cells and HEK Cells. Front Physiol. 2021;12 :736585. doi: 10.3389/fphys.2021.736585. PubMed PMID:34737711 PubMed Central PMC8562563.
- Rapetti-Mauss, R, Berenguier, C, Allegrini, B, Soriani, O. Interplay Between Ion Channels and the Wnt/β-Catenin Signaling Pathway in Cancers. Front Pharmacol. 2020;11 :525020. doi: 10.3389/fphar.2020.525020. PubMed PMID:33117152 PubMed Central PMC7552962.
- Potier-Cartereau, M, Raoul, W, Weber, G, Mahéo, K, Rapetti-Mauss, R, Gueguinou, M et al.. Potassium and Calcium Channel Complexes as Novel Targets for Cancer Research. Rev Physiol Biochem Pharmacol. 2022;183 :157-176. doi: 10.1007/112_2020_24. PubMed PMID:32767122 .
- Martin, N, Soriani, O, Bernard, D. Cardiac Glycosides as Senolytic Compounds. Trends Mol Med. 2020;26 (3):243-245. doi: 10.1016/j.molmed.2020.01.001. PubMed PMID:31983612 .
- Soriani, O, Kourrich, S. The Sigma-1 Receptor: When Adaptive Regulation of Cell Electrical Activity Contributes to Stimulant Addiction and Cancer. Front Neurosci. 2019;13 :1186. doi: 10.3389/fnins.2019.01186. PubMed PMID:31780884 PubMed Central PMC6861184.
- Rapetti-Mauss, R, Borgese, F, Harvey, BJ, Soriani, O. [KCNQ1: a new regulator of the epithelio-mesenchymal transition in colorectal cancers]. Med Sci (Paris). 2018;34 (1):21-24. doi: 10.1051/medsci/20183401006. PubMed PMID:29384089 .
- Rapetti-Mauss, R, Picard, V, Guitton, C, Ghazal, K, Proulle, V, Badens, C et al.. Red blood cell Gardos channel (KCNN4): the essential determinant of erythrocyte dehydration in hereditary xerocytosis. Haematologica. 2017;102 (10):e415-e418. doi: 10.3324/haematol.2017.171389. PubMed PMID:28619848 PubMed Central PMC5622875.
- Rapetti-Mauss, R, Bustos, V, Thomas, W, McBryan, J, Harvey, H, Lajczak, N et al.. Bidirectional KCNQ1:β-catenin interaction drives colorectal cancer cell differentiation. Proc Natl Acad Sci U S A. 2017;114 (16):4159-4164. doi: 10.1073/pnas.1702913114. PubMed PMID:28373572 PubMed Central PMC5402468.
- Soriani, O, Rapetti-Mauss, R. Sigma 1 Receptor and Ion Channel Dynamics in Cancer. Adv Exp Med Biol. 2017;964 :63-77. doi: 10.1007/978-3-319-50174-1_6. PubMed PMID:28315265 .
- Rapetti-Mauss, R, Soriani, O, Vinti, H, Badens, C, Guizouarn, H. Senicapoc: a potent candidate for the treatment of a subset of hereditary xerocytosis caused by mutations in the Gardos channel. Haematologica. 2016;101 (11):e431-e435. doi: 10.3324/haematol.2016.149104. PubMed PMID:27443288 PubMed Central PMC5394861.
- Guéguinou, M, Harnois, T, Crottes, D, Uguen, A, Deliot, N, Gambade, A et al.. SK3/TRPC1/Orai1 complex regulates SOCE-dependent colon cancer cell migration: a novel opportunity to modulate anti-EGFR mAb action by the alkyl-lipid Ohmline. Oncotarget. 2016;7 (24):36168-36184. doi: 10.18632/oncotarget.8786. PubMed PMID:27102434 PubMed Central PMC5094991.
- Crottès, D, Félix, R, Meley, D, Chadet, S, Herr, F, Audiger, C et al.. Immature human dendritic cells enhance their migration through KCa3.1 channel activation. Cell Calcium. 2016;59 (4):198-207. doi: 10.1016/j.ceca.2016.02.008. PubMed PMID:27020659 .
- Crottès, D, Rapetti-Mauss, R, Alcaraz-Perez, F, Tichet, M, Gariano, G, Martial, S et al.. SIGMAR1 Regulates Membrane Electrical Activity in Response to Extracellular Matrix Stimulation to Drive Cancer Cell Invasiveness. Cancer Res. 2016;76 (3):607-18. doi: 10.1158/0008-5472.CAN-15-1465. PubMed PMID:26645564 .
- Rapetti-Mauss, R, Lacoste, C, Picard, V, Guitton, C, Lombard, E, Loosveld, M et al.. A mutation in the Gardos channel is associated with hereditary xerocytosis. Blood. 2015;126 (11):1273-80. doi: 10.1182/blood-2015-04-642496. PubMed PMID:26148990 .
2013 - Prime d'Excellence Scientifique - Université Nice Sophia Antipolis

Raphaël Rapetti-Mauss: Médaille de Bronze du CNRS 2023
Read More
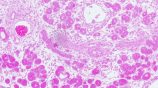
SK2 channels: a signaling hub in pancreatic cancer
Read More
iBV - Institut de Biologie Valrose
"Sciences Naturelles"
Université Nice Sophia Antipolis
Faculté des Sciences
Parc Valrose
06108 Nice cedex 2
