
Olivier HUMBERT
PET/CT imaging and Artificial Intelligence for precision Oncology
Main interests
- PET and CT imaging
- Imaging of tumor response to immuno-chemotherapy
- Implementation of AI models for real-world precision oncology
Scientific Questions
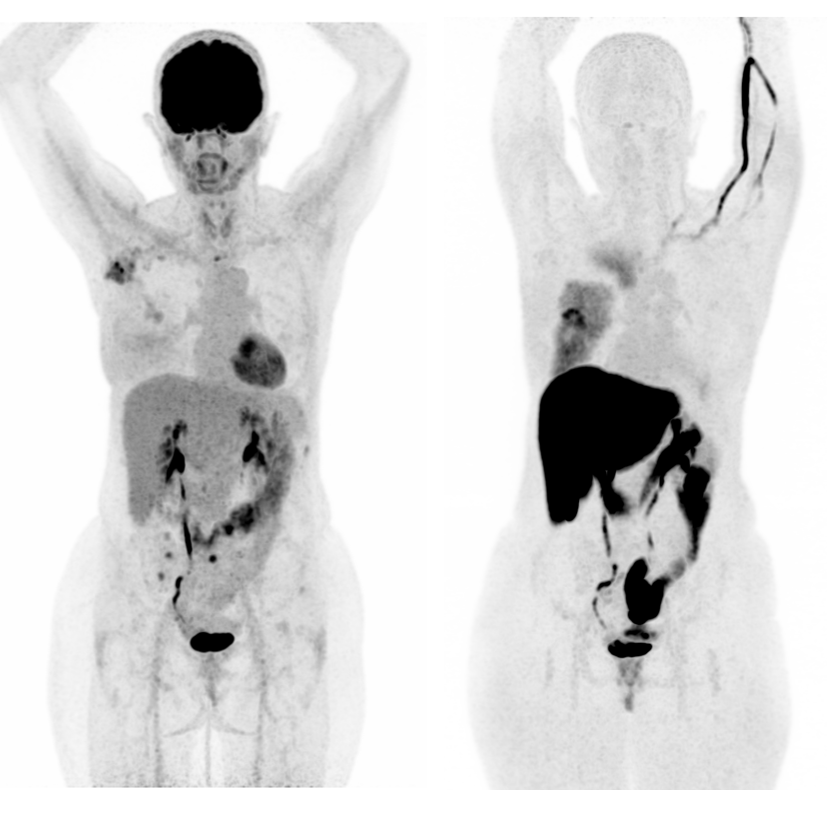
Recent therapeutic advances have significantly improved outcomes for patients with metastatic cancer. However, therapeutic resistance remains a major challenge, highlighting the urgent need for novel predictive biomarkers to better guide treatment strategies.
In parallel, assessing tumor response using whole-body imaging modalities has become increasingly complex, especially in the era of immunotherapy. Atypical response patterns, including pseudo-progression and dissociated responses, limit the reliability of current radiological criteria of tumor response such as RECIST 1.1 or PERCIST 1.0.
To address these challenges, there is a critical need to develop new imaging biomarkers and response assessment frameworks. In this context, advanced analytical approaches, particularly those based on artificial intelligence, offer promising opportunities to integrate complex imaging patterns of response and clinical data, refine the current response criteria, enable more accurate monitoring of treatment efficacy, and finally improve patient’s outcome prediction.
Our Strategy
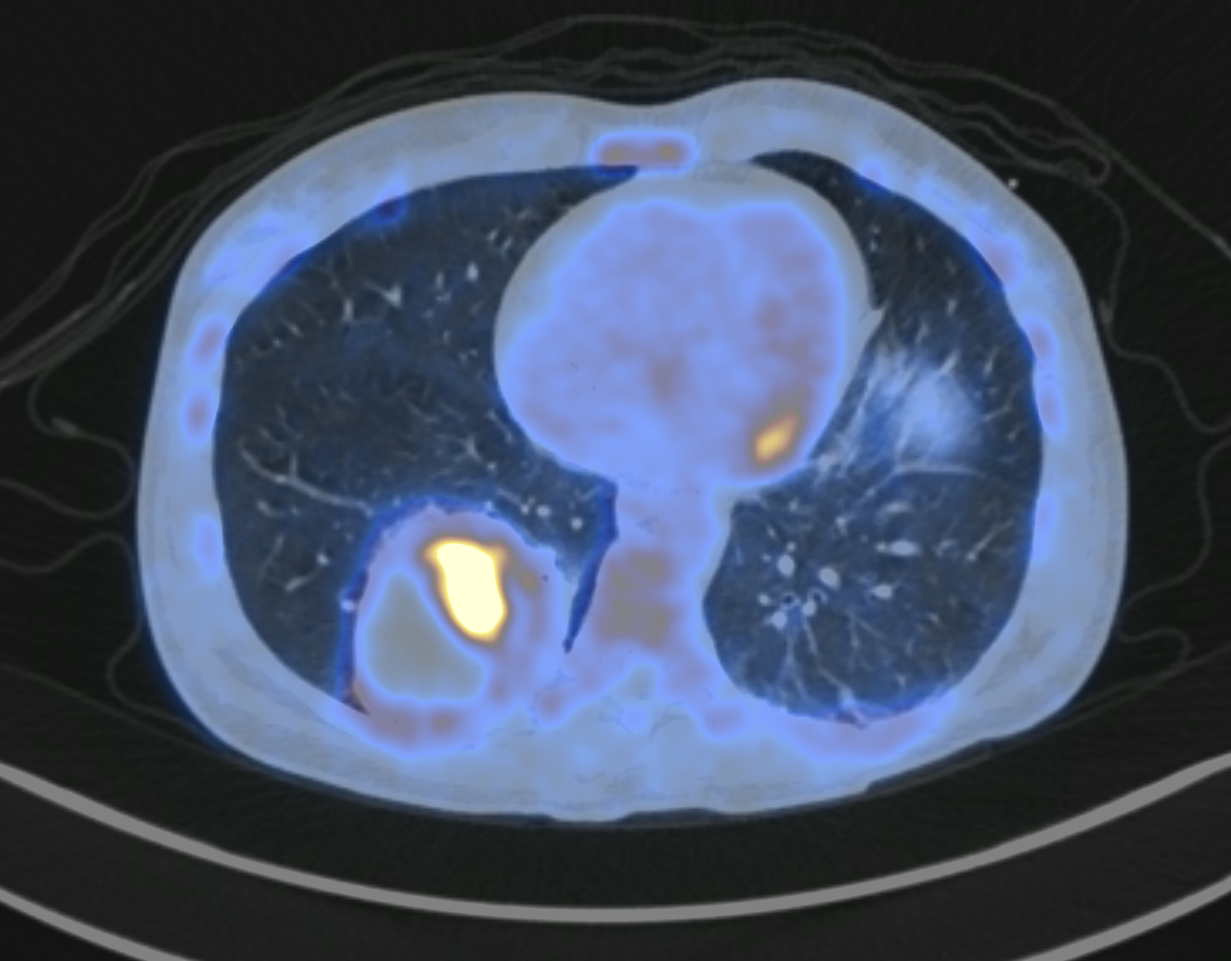
Among the main issues for development of AI algorithms is the availability of large, curated and multicentric training medical data, stored in a unique and centralized repository. To tackle this problem, work is being carried out on so-called federated learning technologies (PRT-K 2022 funding). The FEDERATED-PET project aims at establishing the first French Federated Learning (FL) initiative for the collaborative Deep Learning analysis of 3D PET/CT images and clinical data, leveraging on state-of-the-art FL platforms (FedBio-Med, INRIA) and an important network of hospitals. FL provides will create a unique opportunity to collaboratively learn a shared model across multiple hospitals, solving the crucial problem of medical data sharing and exchange.
The ultimate goal of our strategy is the clinical validation and translation of these AI models into real-world settings. To achieve this, we maintain very close collaboration with hospital partners (Centre Antoine Lacassagne, Unicancer Fedrartion...) and medical imaging companies, ensuring access to high-quality multicentric data for model training and validation. These partnerships aim to accelerate deployment of our models and deliver tangible benefits for patient care in oncology.
Research Aims
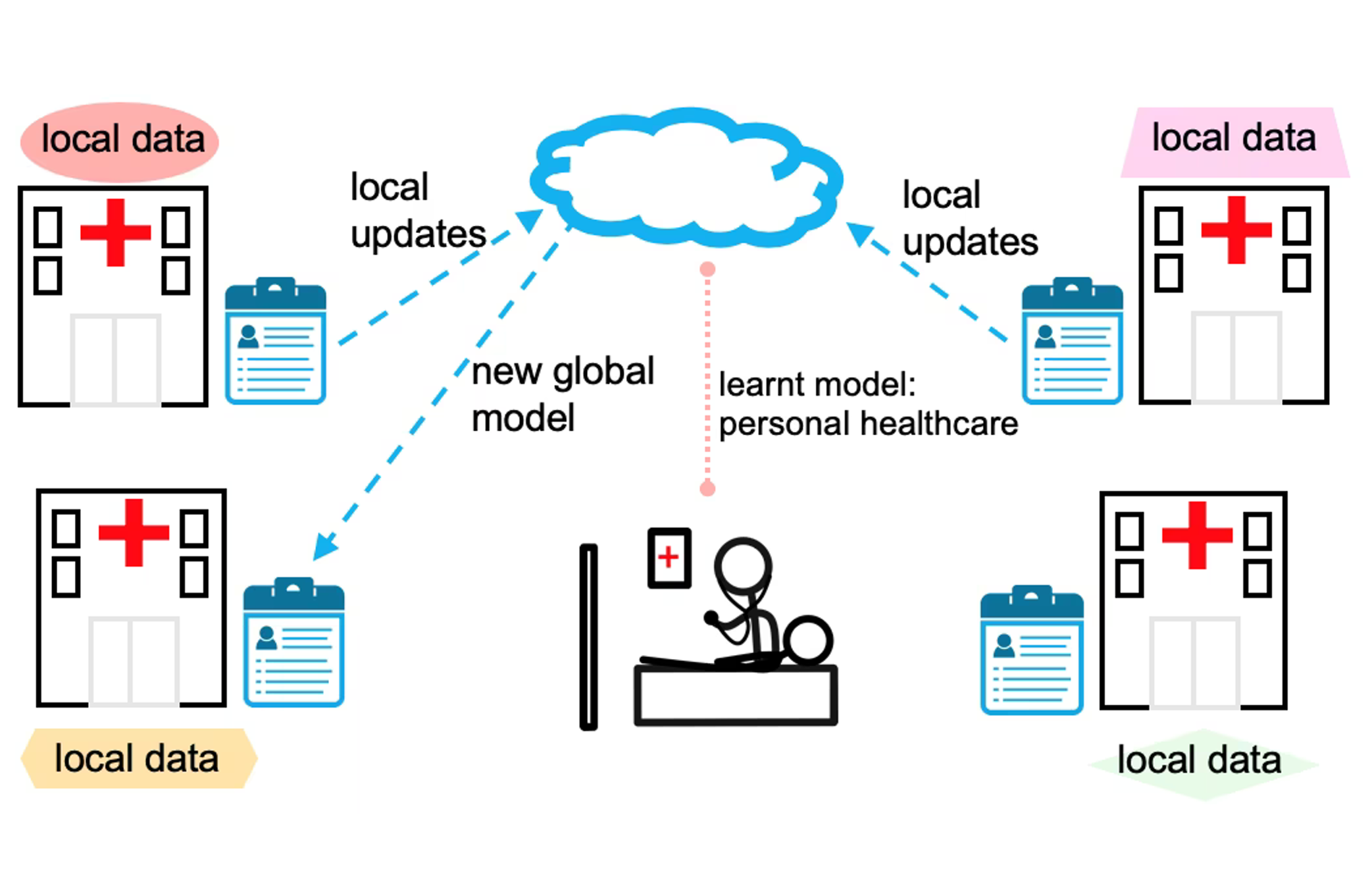
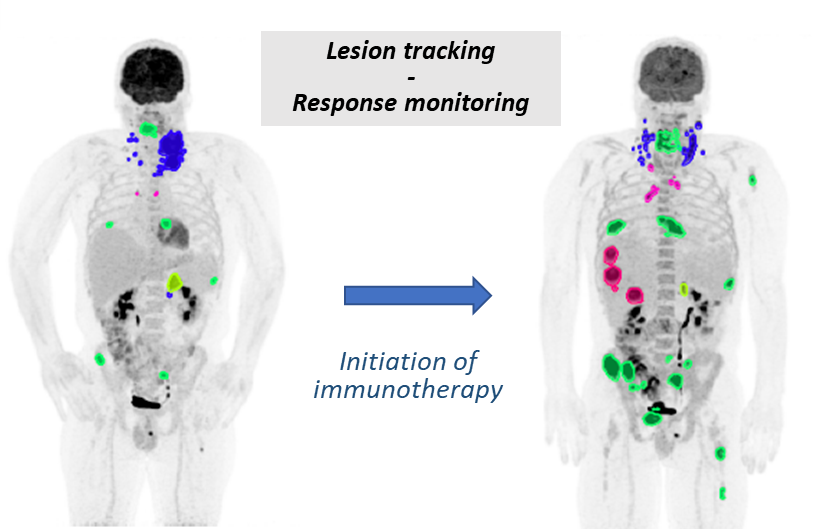
Our goal is to redefine the assessment of therapeutic response in metastatic cancer. We seek to move beyond conventional, lesion-level evaluation toward a patient-level, dynamic, and predictive framework that comprehensively captures tumor heterogeneity, temporal evolution, and interactions with the immune system and patients’ organ. Specifically, we aim to:
- develop novel AI architectures capable of integrating clinical, imaging and biological data
- implement longitudinal models to detect early and atypical imaging response patterns, including pseudo-progression and dissociated responses, which are undetectable with current response evaluation criteria in solid tumors
- leverage federated learning technologies to enable secure, multicentric access to medical data for robust model training
- achieve real-world clinical validation of AI models. Beyond technical performance, we will therefore evaluate their consistency across diverse clinical environments and their practical added value in routine care.
Researchers
 CHARDIN David - +33 R
CHARDIN David - +33 R
PreDocs
 SCHMUTZ Hugo - +33 R
SCHMUTZ Hugo - +33 R COMTE Victor - +33 R
COMTE Victor - +33 R MASSE Mathilde - +33 R
MASSE Mathilde - +33 R MOHEBI Mobin - +33 R
MOHEBI Mobin - +33 R NGUYEN Huyen-Trang - +33 R
NGUYEN Huyen-Trang - +33 R FORONI Giulia - +33 R
FORONI Giulia - +33 R
Recent publications
- Gambiez, M, Palard-Novello, X, Guéry, C, Marchal, E, Dercle, L, Morland, D et al.. Multicenter evaluation of commercial AI for [18F]FDG PET/CT lesion detection. Ann Nucl Med. 2026; :. doi: 10.1007/s12149-026-02199-9. PubMed PMID:41910857 .
- Darcourt, J, Koulibaly, PM, Schiazza, A, Comte, V, Humbert, O. Assessment of striatal and extrastriatal [18F]fluoroDOPA uptake by a Silicon Photomultiplier PET camera in Parkinson's disease. EJNMMI Res. 2026; :. doi: 10.1186/s13550-026-01400-4. PubMed PMID:41781550 .
- Masse, M, Bailleux, C, Creisson, A, Humbert, O. [Molecular imaging and radioligand in breast cancer]. Bull Cancer. 2025;112 (7-8):702-713. doi: 10.1016/j.bulcan.2025.02.014. PubMed PMID:40300962 .
- Ahrari, S, Zaragori, T, Zinsz, A, Hossu, G, Oster, J, Allard, B et al.. Clinical impact of an explainable machine learning with amino acid PET imaging: application to the diagnosis of aggressive glioma. Eur J Nucl Med Mol Imaging. 2025;52 (6):1989-2001. doi: 10.1007/s00259-024-07053-6. PubMed PMID:39821662 .
- Le, TK, Comte, V, Darcourt, J, Razzouk-Cadet, M, Rollet, AC, Orlhac, F et al.. Performance and Clinical Impact of Radiomics and 3D-CNN Models for the Diagnosis of Neurodegenerative Parkinsonian Syndromes on 18 F-FDOPA PET. Clin Nucl Med. 2024;49 (10):924-930. doi: 10.1097/RLU.0000000000005392. PubMed PMID:39104036 .
- Janho Dit Hreich, S, Humbert, O, Pacé-Loscos, T, Schiappa, R, Juhel, T, Ilié, M et al.. Plasmatic Inactive IL-18 Predicts a Worse Overall Survival for Advanced Non-Small-Cell Lung Cancer with Early Metabolic Progression after Immunotherapy Initiation. Cancers (Basel). 2024;16 (12):. doi: 10.3390/cancers16122226. PubMed PMID:38927931 PubMed Central PMC11202099.
- Masse, M, Chardin, D, Tricarico, P, Ferrari, V, Martin, N, Otto, J et al.. [18F]FDG-PET/CT atypical response patterns to immunotherapy in non-small cell lung cancer patients: long term prognosis assessment and clinical management proposal. Eur J Nucl Med Mol Imaging. 2024;51 (12):3696-3708. doi: 10.1007/s00259-024-06794-8. PubMed PMID:38896129 PubMed Central PMC11457717.
- Tricarico, P, Chardin, D, Martin, N, Contu, S, Hugonnet, F, Otto, J et al.. Total metabolic tumor volume on 18F-FDG PET/CT is a game-changer for patients with metastatic lung cancer treated with immunotherapy. J Immunother Cancer. 2024;12 (4):. doi: 10.1136/jitc-2023-007628. PubMed PMID:38649279 PubMed Central PMC11043703.
- Bailleux, C, Chardin, D, Guigonis, JM, Ferrero, JM, Chateau, Y, Humbert, O et al.. Survival analysis of patient groups defined by unsupervised machine learning clustering methods based on patient metabolomic data. Comput Struct Biotechnol J. 2023;21 :5136-5143. doi: 10.1016/j.csbj.2023.10.033. PubMed PMID:37920813 PubMed Central PMC10618114.
- Bailleux, C, Chardin, D, Gal, J, Guigonis, JM, Lindenthal, S, Graslin, F et al.. Metabolomic Signatures of Scarff-Bloom-Richardson (SBR) Grade in Non-Metastatic Breast Cancer. Cancers (Basel). 2023;15 (7):. doi: 10.3390/cancers15071941. PubMed PMID:37046602 PubMed Central PMC10093598.
- Lévi-Strauss, T, Tortorici, B, Lopez, O, Viau, P, Ouizeman, DJ, Schall, B et al.. Radiomics, a Promising New Discipline: Example of Hepatocellular Carcinoma. Diagnostics (Basel). 2023;13 (7):. doi: 10.3390/diagnostics13071303. PubMed PMID:37046521 PubMed Central PMC10093101.
- Chardin, D, Gille, C, Pourcher, T, Humbert, O, Barlaud, M. Learning a confidence score and the latent space of a new supervised autoencoder for diagnosis and prognosis in clinical metabolomic studies. BMC Bioinformatics. 2022;23 (1):361. doi: 10.1186/s12859-022-04900-x. PubMed PMID:36050631 PubMed Central PMC9434875.
- Khatir, W, Humbert, O, Benzaquen, J, Bontoux, C, Neels, J, Berland, L et al.. Identification of a circulating immunological signature predictive of response to immune checkpoint inhibitors in patients with advanced non-small cell lung cancer. Clin Transl Med. 2022;12 (8):e1018. doi: 10.1002/ctm2.1018. PubMed PMID:35994416 PubMed Central PMC9394752.
- Comte, V, Schmutz, H, Chardin, D, Orlhac, F, Darcourt, J, Humbert, O et al.. Development and validation of a radiomic model for the diagnosis of dopaminergic denervation on [18F]FDOPA PET/CT. Eur J Nucl Med Mol Imaging. 2022;49 (11):3787-3796. doi: 10.1007/s00259-022-05816-7. PubMed PMID:35567626 PubMed Central PMC9399031.
- Humbert, O, Bauckneht, M, Gal, J, Paquet, M, Chardin, D, Rener, D et al.. Prognostic value of immunotherapy-induced organ inflammation assessed on 18FDG PET in patients with metastatic non-small cell lung cancer. Eur J Nucl Med Mol Imaging. 2022;49 (11):3878-3891. doi: 10.1007/s00259-022-05788-8. PubMed PMID:35562529 PubMed Central PMC9399195.
- Chardin, D, Humbert, O, Bailleux, C, Burel-Vandenbos, F, Rigau, V, Pourcher, T et al.. Primal-dual for classification with rejection (PD-CR): a novel method for classification and feature selection-an application in metabolomics studies. BMC Bioinformatics. 2021;22 (1):594. doi: 10.1186/s12859-021-04478-w. PubMed PMID:34911437 PubMed Central PMC8672607.
- Humbert, O, Chardin, D. Dissociated Response in Metastatic Cancer: An Atypical Pattern Brought Into the Spotlight With Immunotherapy. Front Oncol. 2020;10 :566297. doi: 10.3389/fonc.2020.566297. PubMed PMID:33072599 PubMed Central PMC7531255.
- Chardin, D, Paquet, M, Schiappa, R, Darcourt, J, Bailleux, C, Poudenx, M et al.. Baseline metabolic tumor volume as a strong predictive and prognostic biomarker in patients with non-small cell lung cancer treated with PD1 inhibitors: a prospective study. J Immunother Cancer. 2020;8 (2):. doi: 10.1136/jitc-2020-000645. PubMed PMID:32709713 PubMed Central PMC7380842.
- Gal, J, Bailleux, C, Chardin, D, Pourcher, T, Gilhodes, J, Jing, L et al.. Comparison of unsupervised machine-learning methods to identify metabolomic signatures in patients with localized breast cancer. Comput Struct Biotechnol J. 2020;18 :1509-1524. doi: 10.1016/j.csbj.2020.05.021. PubMed PMID:32637048 PubMed Central PMC7327012.
- Hoog, C, Koulibaly, PM, Dejean, C, Desdoits, T, Humbert, O, Barranger, E et al.. Comparison of 3 γ-probes for simultaneous iodine-125-seed and technetium-99m breast cancer surgery: NEMA standard characterisation with extended processing. EJNMMI Phys. 2020;7 (1):37. doi: 10.1186/s40658-020-00299-7. PubMed PMID:32504305 PubMed Central PMC7275111.
2024 - Prix Unicancer de l'innovation 2024, en collaboration avec Marco Lorenzi (chercheur Inria)
2011 - Award-winner at the Faculty of Medicine, Dijon, France (Prix de these).
iBV - Institut de Biologie Valrose
"Tour Pasteur"
Université Nice Sophia Antipolis
Faculté de médecine
28 Avenue de Valombrose
06189 Nice cedex 2
